Biologic Bloodhounds Tracking the Paths in RA Pathology
Two new strategies for breaking through the tangle of pathways to identify those most important to rheumatoid arthritis treatment converge on a shared goal: personalized medicine.
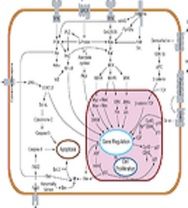
Rheumatic diseases are as complex in their pathology as in the many patterns of symptoms that patients bring to the clinic, seeking relief. How will it ever be possible to sort out all the genes that control the pathologic pathways that create conditions like rheumatoid arthritis (RA), which must happen if "personalized medicine" is to become a reality, someday?
New tactics that break through the complicated tangle of pathways and identify those that matter most to RA treatment were the subject of presentations at the recent American College of Rheumatology annual meeting in San Diego from two research teams, working independently and converging on the same goal.
Gary Firestein MD of the UCSD School of Medicine in San Diego and his team have compiled information from several large databases to sort out patterns in RA pathogenic pathways.1 Researchers in the laboratory of Robert Plenge MD at Harvard Medical School’s Brigham & Women’s Hospital in Boston told how, by linking susceptibility genes to electronic medical records, they have been able to identify variants linked to protection against RA and to adverse events in one pathway important to drug development.2
The UCSD research, funded by the Rheumatology Research Foundation, works at a meta-level above either of the usual methods of associating genes with disorders: (1) the one-on-one analyses that detect relationships between disease traits and individual variants in gene fragments called single nucleotide polymorphisms (SNPs), or (2) the newer genome-wide association studies (GWAS) that correlate physical traits with variations at thousands of locations in the genome at once.3,4
A problem with the latter approach is that “it can identify thousands of genes as being implicated in a condition,” explained John Whitaker PhD, a postdoc who worked with Firestein and biochemist Wei Wang PhD at UCSD on this research. “We know these genes are important, but prioritizing the list of genes can be tricky.”
The researchers addressed this issue by identifying a subset of RA-related genes common to three different data sets:
• DNA sequence variants from the NCBI GWAS4, an online database of published SNPs maintained by the US National Center for Biotechnology Information;5
• Data on gene expression housed in the NIH Gene Expression Omnibus database;
• Their own information about DNA methylation (a process that affects gene expression and therefore sometimes pathogenesis), both from normal cells and from a set of genes identified by the team as involved in RA and osteoarthritis.
“The idea is that each individual data set is noisy. Integrating them is likely to enhance the signal-to-noise ratio and thus to identify genes critical for RA pathogenesis,“ explained Wei Wang MD, senior investigator of the study.
Then they matched the subset of RA genes they had identified against the Kyoto Encyclopedia of Genes and Genomes (KEGG), an online database that summarizes the role of genes in biochemical pathways and diseases.7,8 This has pointed to seven genes and at least four pathways whose regulation may be abnormal in RA. These pathways include:
• Cell adhesion molecules
• Cytokine-cytokine receptor interactions
• Antigen processing and presentation, and
• JAK-STAT signaling.
“The top-ranked pathways are closely related to RA and autoimmune disease,” said Wang. “This work exemplifies the importance of integrated analysis in studying disease mechanisms and identification of new drug targets in the genomic era.”
A different kind of integration informs the work of Robert Plenge MD and a team at Brigham and Women’s Hospital of Harvard Medical School, who have linked RA susceptibility genes to clinical diagnoses using electronic medical records (EMRs).8 Because disease-associated genes can mimic pharmacological effects, they propose, linking genetic data to records in this way can provide important insights for drug development.
The team genotyped 1,118 patients with RA and an equal number without RA, searching for informative variants in exons (protein-coding sections) of genes related to RA. The search focused their attention on the gene TYK2, which codes for a Janus kinase, an enzyme in the JAK-STAT cytokine signaling pathway that is the target of many new RA drugs including tofacitinib.
Deeper analysis with an even larger dataset (9,372 RA cases and 18,868 controls) revealed some TYK2 variants that are associated with clinical diagnoses related to adverse drug events, especially infections, as well as several loss-of-function variants that appear to protect against RA by disabling the enzyme. Studies by other teams have linked these protective variants with reduced TYK2 kinase activity, including impaired pro-inflammatory cytokine signaling, according to Dorothe Diogo PhD, first author of the study.
“The observation of independent disease-associated variants unequivocally highlights the role of TYK2 in RA disease susceptibility,” declared Diogo, “and provides an accumulation of evidence that a drug that mimics the effect of TYK2 alleles may be effective at treating RA.”
The take-home message, she added: Integrating human genetics and real-world data from EMRs can help ascertain efficacy and adverse effects of new drugs before they enter human trials – an especially valuable approach for complex diseases like RA.
References:
1. Whitaker, J, Wang W, Firestein G. Key Rheumatoid Arthritis-Associated Pathogenic Pathways Revealed by Integrative Analysis of RA Omics Datasets. Abstract presented at the ACR Conference, San Diego. Oct. 30, 2013. 2. Plenge R, Diogo D; Liao K, et al.Potential Of Integrating Human Genetics and Electronic Medical Records For Drug Discovery: The Example Of TYK2 and Rheumatoid Arthritis. Abstract presented at the ACR Annual Conference, San Diego. Oct 30, 2013. 3. C.M. Lewis. Genetic association studies: Design, analysis and interpretation.Briefings in Bioinformatics. (2002) 3(2):146–153. 4. B.E. Stranger, E. A. Stahl, and T. Raj. Progress and Promise of Genome-Wide Association Studies for Human Complex Trait Genetics.Genetics (2011) 187:367-383. 5. Hindorff LA, MacArthur J (European Bioinformatics Institute), Morales J (European Bioinformatics Institute), Junkins HA, et al. A Catalog of Published Genome-Wide Association Studies. Available online at: www.genome.gov/gwastudies. 6. Hindorff LA, Sethupathy P, Junkins HA, et al.. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc Natl Acad Sci USA. (2009) 106: 9362-9367. 7. Kyoto Encyclopedia of Genes and Genomes. Available online at http://www.genome.jp/kegg/ 8. Kyoto Encyclopedia of Genes and Genomes: Rheumatoid Arthritis Homo sapiens Pathways.